 So in a situation in which battle between PH and pka is the pt is less and if the pka in more than that means that more poston out side it means molecule going to bind with proton that in they will going, proteinated jej poeton Badded to the pepride (PH/pkal) If the pH h more and if the pka is less than that means that more of ions in the envorment it moars molecule to de protenated. This project is in no way not affiliated, sponsored, or otherwise endorsedīy the original Peptides authors.The structure of peptide is given by plaz 10.5 Ho O H H ha Pk 9.6 pka=2.0 HOT Pe: 10.5 are they going Pka = 6.0 peptide going to take a proton from the solution or going to donate a proton to the solution which can i figur out with the value from pka. Released under the GNU General Public License v1.0. The terms of the GNU General Public License v2.0. The original R Peptides package was written by Daniel Osorio, This library is provided under the GNU General Public License v3.0. In a simple, easily reproducible situation. Information as you can about the issue, and try to recreate the same bug If you are filing in on a bug, please include as much 0.299333 0.465333 - 0.976667 0.023333 □ Feedback ⚠️ Issue Trackerįound a bug ? Have an enhancement request ? Head over to the GitHub issue DataFrame () > df BLOSUM1 BLOSUM2 BLOSUM3 BLOSUM4. This makes it very easy to create a pandas.DataFrame withĭescriptors for several protein sequences: > seqs = > df = pandas. Use the scriptors method to get a dictionary with every availableĭescriptor. Will return a dedicated named tuple: > peptide. Methods that return more than one scalar value (for instance, Peptide.blosum_indices) Then use the appropriate methods to compute the descriptors you want: > peptide. Peptide ( "MLKKRFLGALAVATLLTLSFGTPVMAQSGSAVFTNEGVTPFAISYPGGGT" ) Start by creating a Peptide object from a protein sequence: > import peptides > peptide = peptides. Which hosts universal wheels that can be installed with pip: $ pip install peptidesĬan be found in the online documentation, Install the peptides package directly from PyPi (like numpy.sum or numpy.take) will be used, otherwise a fallbackįrom the standard library is available. If numpy can be imported, relevant functions Taken in a lookup table, so computing them can be done in a vectorized manner. Most of the descriptors for a protein sequence are simple averages of values Reach out and open a feature request on the issue tracker If this library is missing a useful statistic or descriptor, feel free to Biological properties: structural class prediction.Physicochemical properties: aliphatic index, instability index, theoretical net charge, isoelectric point, molecular weight (with isotope labelling support).Sequence profiles: hydrophobicity, hydrophobic moment, membrane position.  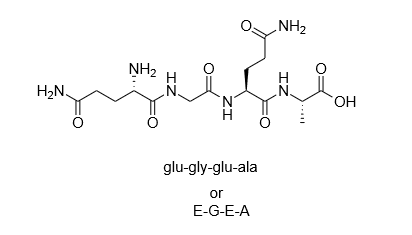  
QSAR descriptors: BLOSUM indices, Cruciani properties, FASGAI vectors, Kidera factors, MS-WHIM scores, PCP descriptors, ProtFP descriptors, Sneath vectors, ST-scales, T-scales, VHSE-scales, Z-scales.Peptide statistics: amino acid counts and frequencies.This library has no external dependency and is available for all modern PythonĪ non-exhaustive list of available features: It started as a port of Peptides, the R package written byĮxPASy Protein Identification and Analysis Tools, and Rcpi. Peptides.py is a pure-Python package to compute common descriptors for Physicochemical properties, indices and descriptors for amino-acid sequences. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
